# Bioconductor
library("airway")
library("ComplexHeatmap")
library("DESeq2")
library("org.Hs.eg.db")
library("SummarizedExperiment")
data(airway)
# data visualization
library("circlize")
library("ggplot2")
library("ggtext")
# data wrangling
library("dplyr")
library("magrittr")
library("tidyr")In the RangedSummarizedExperiment object, we can use the rowData command from SummarizedExperiment to look at meta data that has been collected about the reads. Here, I then present only the column names of that rowData for brevity.
[1] "gene_id" "gene_name" "entrezid" "gene_biotype"
[5] "gene_seq_start" "gene_seq_end" "seq_name" "seq_strand"
[9] "seq_coord_system" "symbol" "exonic_length" In the RangedSummarizedExperiment object, we can use the rowData command from SummarizedExperiment to look at meta data that has been collected about the samples. Here, I then present only a subset of that colData for brevity.
DataFrame with 6 rows and 6 columns
SampleName cell dex albut Run avgLength
<factor> <factor> <factor> <factor> <factor> <integer>
SRR1039508 GSM1275862 N61311 untrt untrt SRR1039508 126
SRR1039509 GSM1275863 N61311 trt untrt SRR1039509 126
SRR1039512 GSM1275866 N052611 untrt untrt SRR1039512 126
SRR1039513 GSM1275867 N052611 trt untrt SRR1039513 87
SRR1039516 GSM1275870 N080611 untrt untrt SRR1039516 120
SRR1039517 GSM1275871 N080611 trt untrt SRR1039517 126Toward Gene Expression
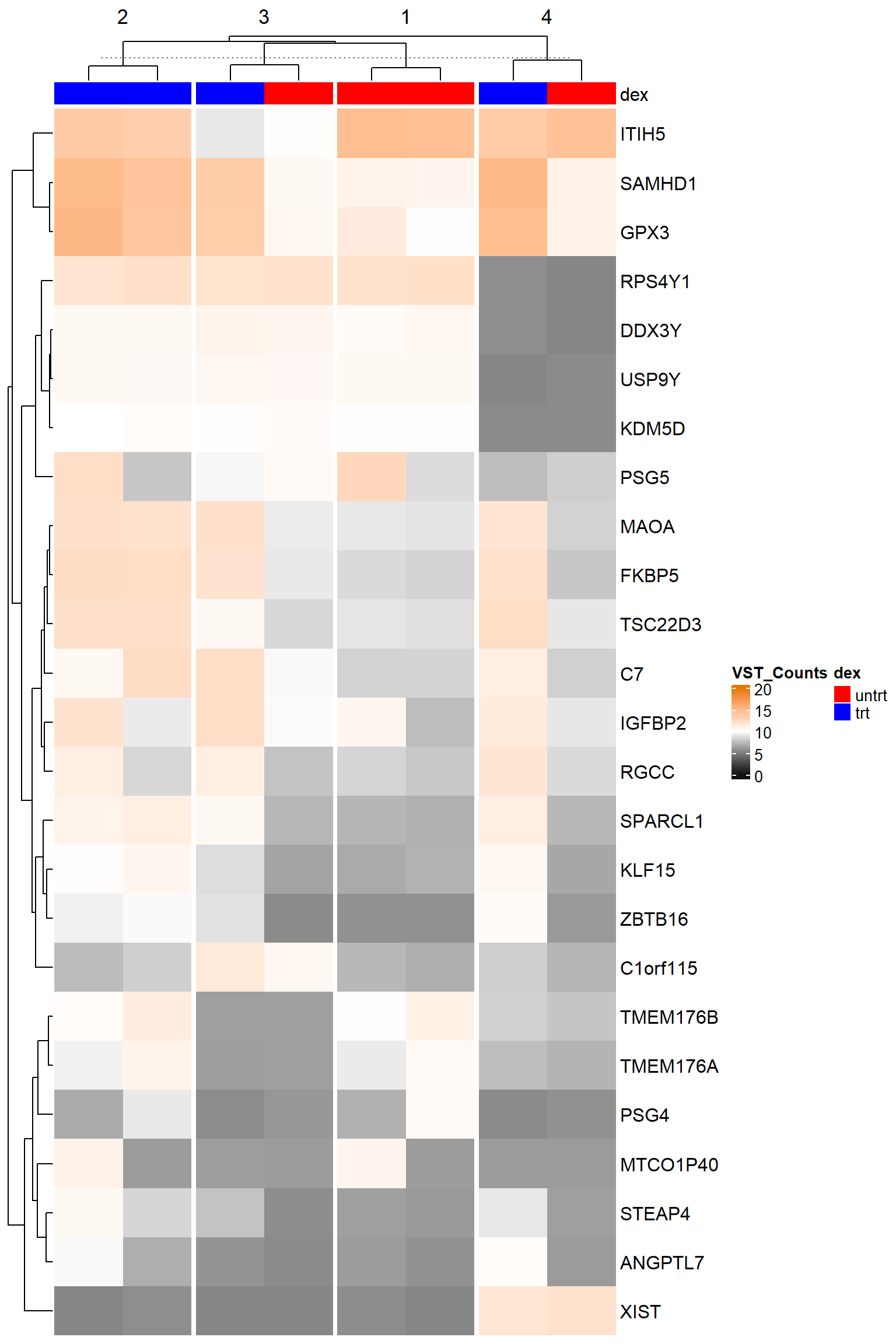
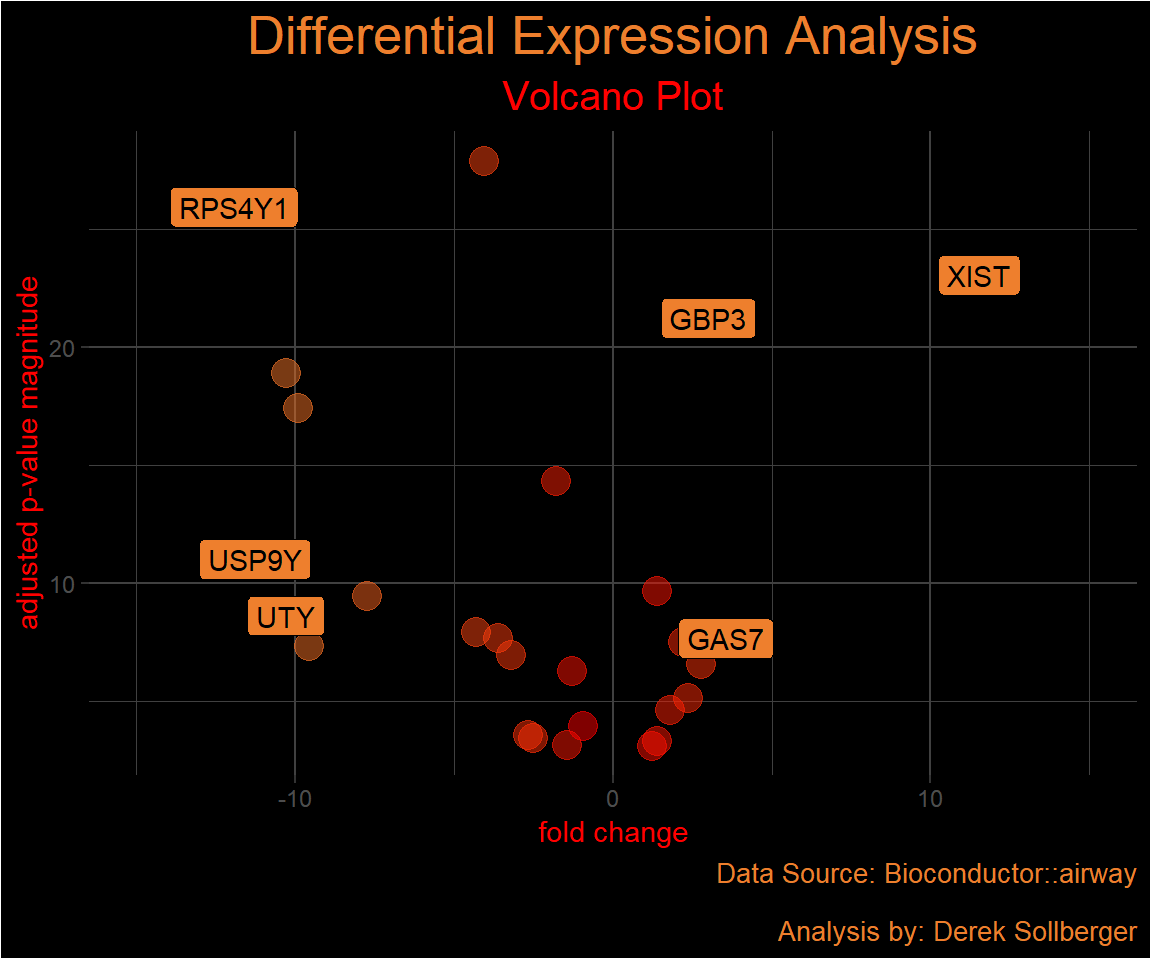
Overall, I want to focus on the genes and the assay of counts of gene expression. Continuing with the EnhancedVolcano documentation, we “annotate the Ensembl gene IDs to gene symbols”.